Evaluating Machine-Learned Inter-Atomic Potentials for a Practical Simulation Workflow
MLIPs are a promising paradigm in atomistic simulation, potentially offering the accuracy of ab-initio methods at the speed of empirical potentials. In this blog post, we give an overview of recent MLIP architectures, followed by an evaluation on a practical CO2 adsorption simulation. We find that as of today these models, though promising, are far from plug-and-play, requiring significant engineering effort to operate within established simulation frameworks, while also failing to produce physically consistent results.
Introduction
Driven by our curiosity about the nanoscopic world, simulating atoms – and by extension molecules – at the quantum level is a highly active area of research going back a century
In this blog post, we aim to provide a structured overview of modern MLIPs, including the key architectural ideas behind them. We then move from theory to practice by examining how these models can be integrated into widely used MD frameworks and what technical friction points arise. Finally, we evaluate practical applicability through targeted benchmarks on inference speed and two realistic case studies (CO₂ diffusion, and CO₂ adsorption in Mg-MOF-74).
Overview of MLIPs
Even though DFT methods heavily improve speed and scaling properties for Ab-Initio MD (AIMD) compared to SOTA coupled cluster methods, they’re still too slow for large scale systems, usually the more interesting ones. One key problem is the fact that DFT approximates the entire wave function, in essence solving the entire system completely, every single time step, which is very costly and scales poorly with system size. But for MD we only care about the forces acting on the atoms, which are derived from the wave function, not the wave function itself. So if there was a way to compute the forces from atomic positions directly, we could skip the highly expensive wave function calculation entirely. MLIPs are trained on a dataset of DFT calculations to learn this mapping as accurately as possible, potentially providing the best of the two worlds: the accuracy of DFT with the speed of cheap empirical potentials.
Explicit MLIPs These take as input, for $N$ atoms, their positions $R \in \mathbb{R}^{N \times 3}$ and species $Z \in \mathbb{Z}^{N}$, and learn the mapping to energy $E \in \mathbb{R}$, forces $F \in \mathbb{R}^{N \times 3}$ and stress $\sigma \in \mathbb{R}^{N \times 6}$ directly. This is considered the straightforward and intuitive approach, but no energy conservation guarantees are offered by it.
\[F_i, E, \sigma_{\alpha \beta} = f_\theta(r_1, r_2, \dots, r_n)\]These MLIPs are also referred to as direct-force predictors.
Implicit MLIPs These take as input, for $N$ atoms, their positions $R \in \mathbb{R}^{N \times 3}$ and species $Z \in \mathbb{Z}^{N}$, and a mapping is learned to the potential energy $E \in \mathbb{R}$ of the system alone:
\[E = f_\theta(r_1, r_2, \dots, r_n)\]The forces $F_i$, the stress $\sigma$, and the strain tensors $\epsilon_{\alpha \beta}$ are then computed by taking the negative gradient of the network with respect to the atomic positions $r_i$ and the strain tensors $\epsilon_{\alpha \beta}$ respectively.
\[\mathbf{F}_i = - \nabla_{\mathbf{r}_i} E \quad\text{and}\quad \sigma_{\alpha \beta} = \frac{1}{V} \frac{\partial E}{\partial \epsilon_{\alpha \beta}}\]They are also called conservative MLIPs.
SchNet (2018)
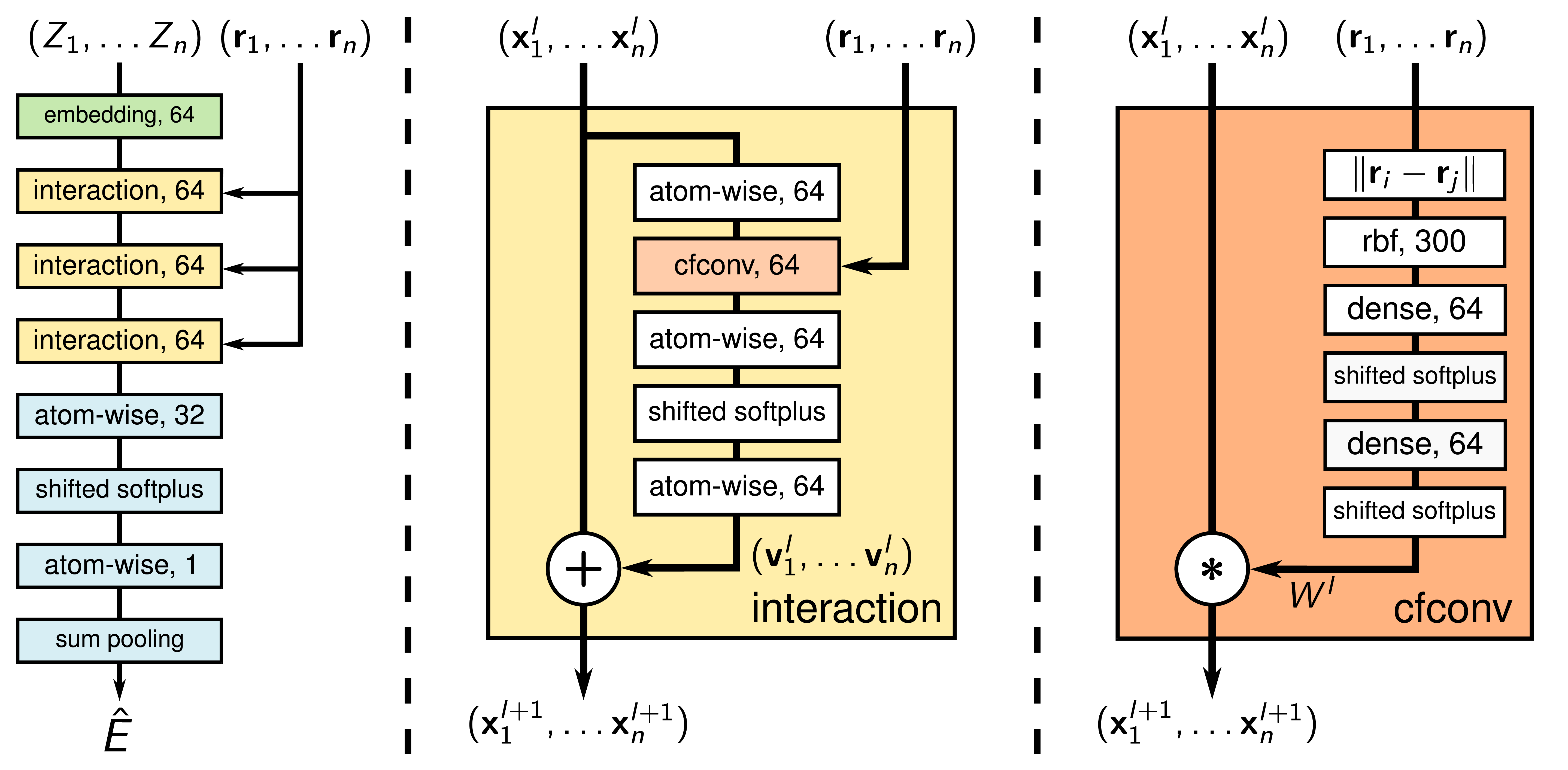
SchNet
Invariance to E(3)
SchNet is invariant to rotations, translations, and atomic indexing, which made it a significant step forward compared to previous models. Translation equivariance and atomic indexing symmetry are the easiest to achieve, as they often come naturally when working with graphs; which is why, going forward, we will only focus on rotational equivariance.
SchNet achieves rotational equivariance by using the invariant distance $d_{ij} = ||r_i - r_j||_2$ between two atoms $i$ and $j$ as input to the model. These distances are then transformed into coefficients of Radial Basis Functions (RBFs),
\(e_k(d_{ij}) = \exp(-\gamma (d_{ij} - \mu_k)^2) \quad\text{with}\quad k \in [1, K],\) where $\mu_k$ are the Gaussian centers, a fix for the high variance of the distance variable slightly resembling Fourier Features
The update step is a simple convolution with ResNet
weighted by a learned function $W(d_{ij})$ of the inter-atomic distances. But newer models no longer take distances alone as model input, due to the fact that they underrepresent atomic systems especially when the model has fewer interaction layers. Only measuring distance can lead to having different atomic systems that result in the same prediction, even though they exert different forces and energies.
Continuous Filter Convolutions
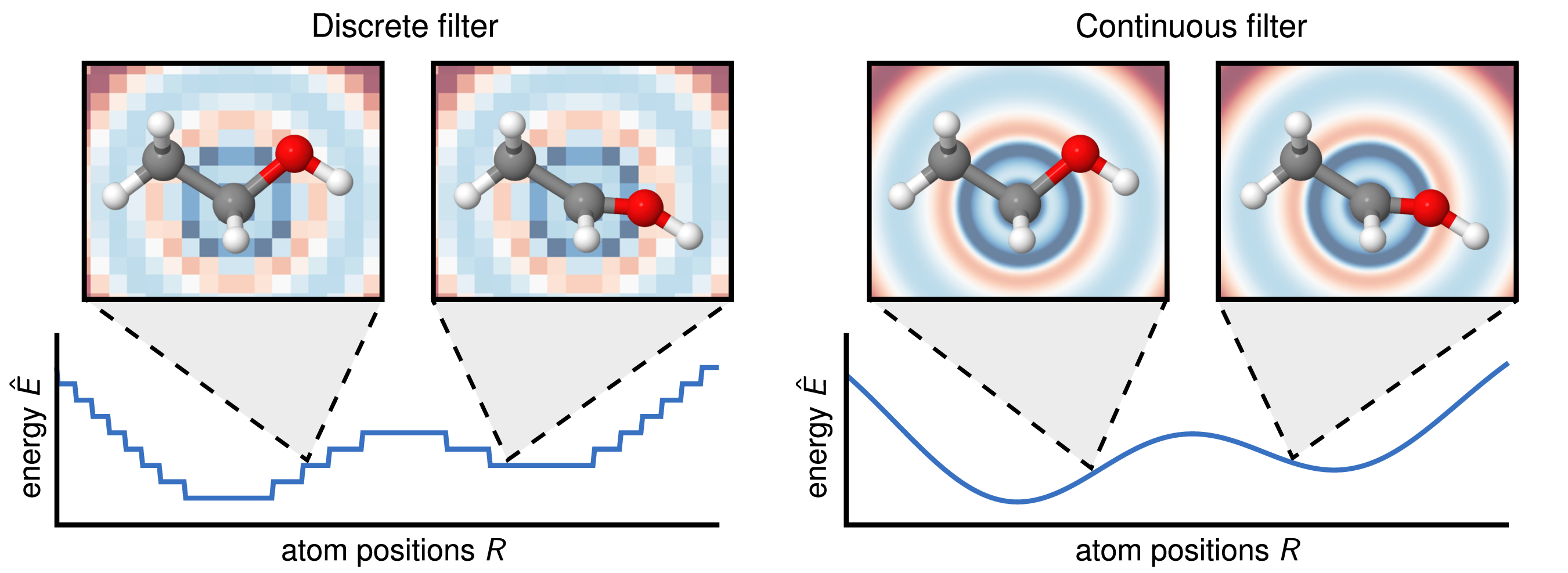
A major challenge for applying convolutions in MLIPs is the irregular and sparse arrangement of neighboring atoms. This contrasts with CNNs for images, where pixels form a regular grid and the discretization is fine enough to make reasonable predictions. For MLIPs, a discrete convolution kernel analogous to those in CNNs results in a poorly discretized, discontinuous PES. SchNet addresses this problem by introducing CFConv layers. Instead of using a discrete kernel matrix, this method learns a continuous kernel function. First, the distance between two atoms $d_{ij}$ is expanded using a set of RBF as shown above. Then, the resulting vector is processed by a small MLP to generate a filter weight $W(d_{ij})$:
\[W(d_{ij}) = \text{MLP}(e_1(d_{ij}), e_2(d_{ij}), \dots, e_K(d_{ij})).\]This technique of generating filter weights from interatomic distances results in a smooth PES, as illustrated below.
NequIP (2022)
NequIP
The key idea is to project the directional information onto spherical harmonics \(Y_m^{(l)}(\hat{r}_{ij})\), which are rotationally equivariant. These spherical harmonics are then multiplied with a learnable radial function \(R(r_{ij})\)
\[S_m^{(l)}(\hat{r}_{ij}) = R(r_{ij}) Y_m^{(l)}(\hat{r}_{ij}),\]to form a convolutional filter similar to SchNet’s CFConv layers. In their experiments they found that using $l=0$ for the spherical harmonics (i.e. rotational invariance) yielded significantly worse results than using $l=1$ or $l=2$, but increasing $l$ further didn’t improve results much more.
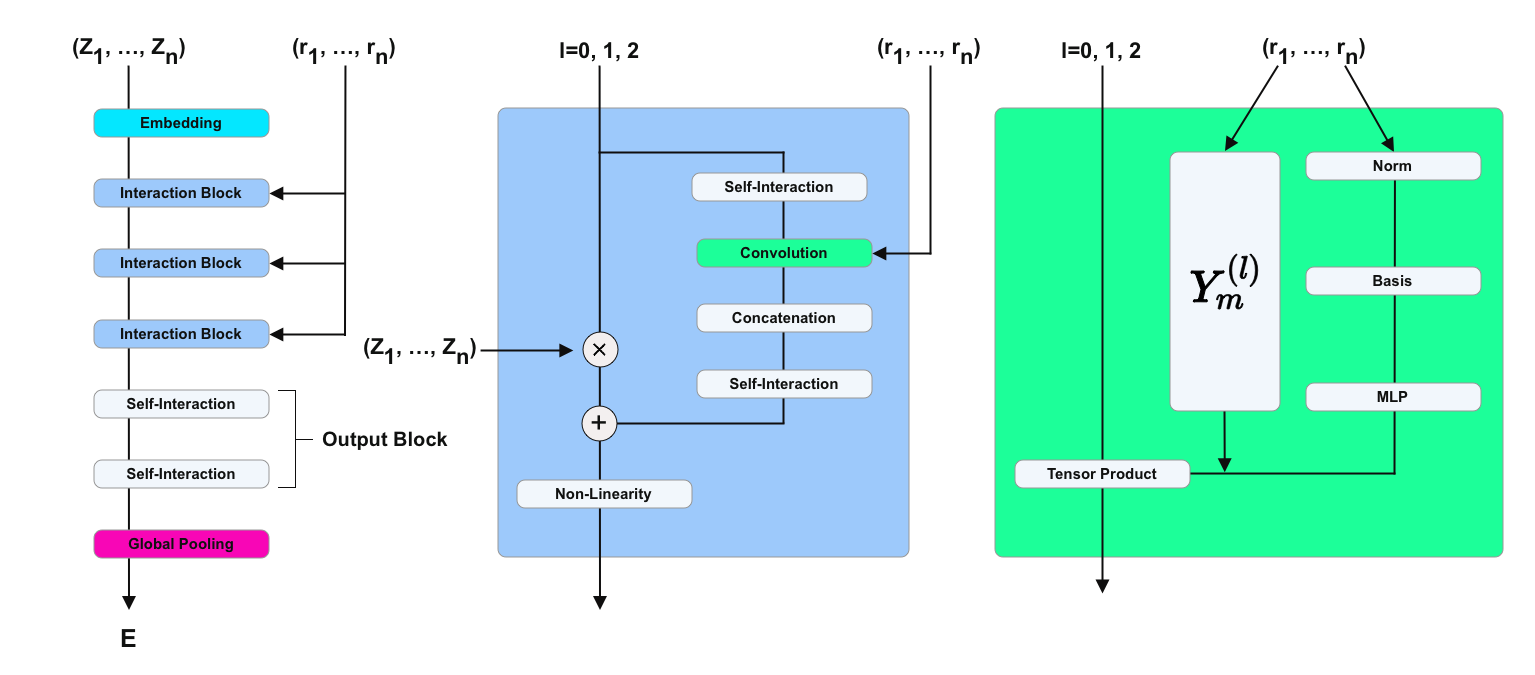
The Clebsch-Gordan tensor product of two SO(3) representations is again a representation of SO(3), which allows them to mix and match features of different orders $l$ freely - a significant improvement over the use of only a limited set of equivariant operations.
Orb-v2 (2024)
The Orb models from Neumann et al.
Orb-v2 is one of the fastest SOTA MLIP models, primarily due to its highly simple architecture and relatively small amount of parameters. It avoids loops and is therefore highly vectorizable, allowing for highly efficient batching and parallelization on modern hardware and uses torch-scatter
eSEN (2025)
The eSEN model from Fu et al. (FAIR)
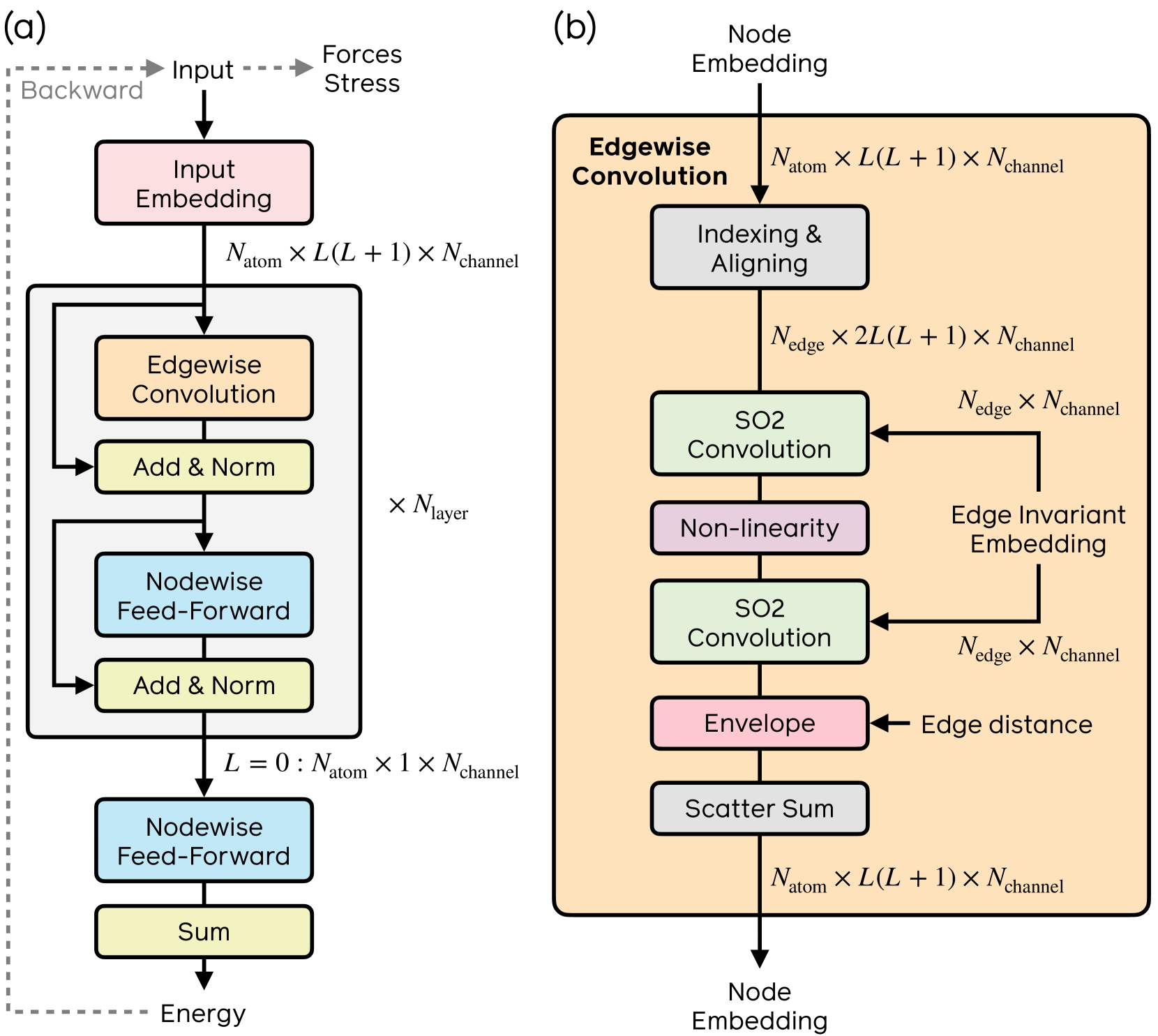
The first key idea is to remove all but one degree of freedom in the message passing step. Generally, with one atom fixed at the center of the coordinate system, the other atom has three degrees of freedom to the first atom left, prompting a full SO(3) equivariant architecture. Zitnick et al.
rotating the result back to the original reference frame. The message network still sees the spherical channels displayed relative to its frame of reference and can therefore still make out relative information of its neighbors – but only has to learn this with one varying degree of freedom, the azimuthal angle $\theta$ around the z-axis, which $f_{\text{message}}$ needs to smoothly convolve over. Spherical harmonics become now become independent of the azimuthal angle $\theta$ and are therefore reduced to circular harmonics, which only depend on the polar angle $\phi$ as shown below.
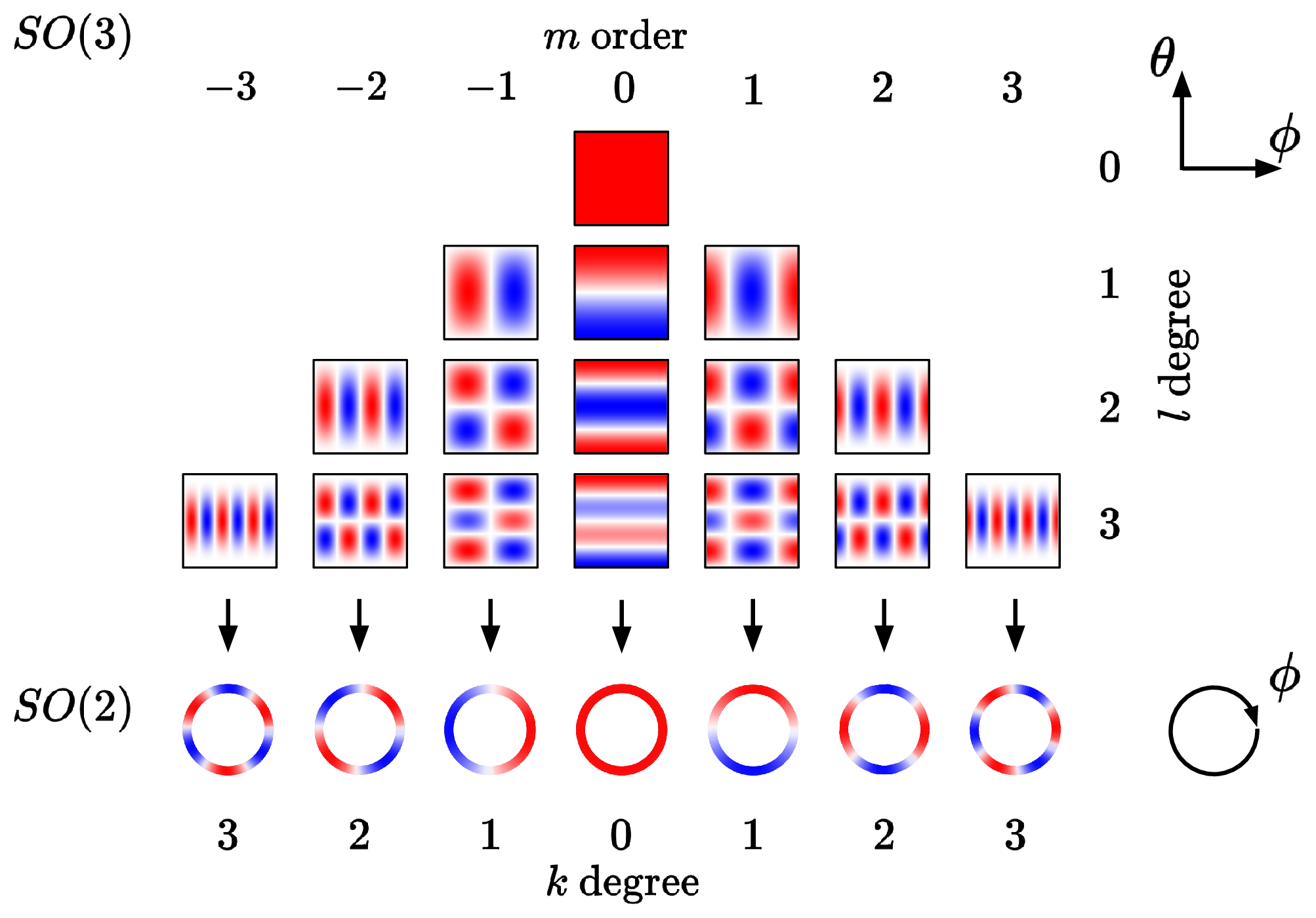
The second key idea is to make use of the fact that a convolution in the spatial domain is equivalent to a multiplication in the frequency domain
And because just like fourier coefficients the spherical harmonics are such frequency components, the convolution of coefficients $x_{l,m,c}$ with filters $h_{l,m,c,l’,c’}$ can be expressed as
\[y_{l,0,c} = \sum_{l'c'} h_{l,0,c,l',c'} x_{l',0,c'}\]where $l$ is the order, $m$ the degree, and $c$ the channel index. But as the coefficients might be complex,
\[\left(\begin{array}{c} y_{l,m,c} \\ y_{l,-m,c} \\ \end{array}\right) = \sum_{l'c'} \left(\begin{array}{cc} h_{l,m,c,l',c'} & -h_{l,-m,c,l',c'} \\ h_{l,-m,c,l',c'} & h_{l,m,c,l',c'} \end{array}\right) \left(\begin{array}{c} x_{l',m,c'} \\ x_{l',-m,c'} \end{array}\right)\]is the general formula they use and provide.
Even though in speed it is ahead of the previous e3nn architectures with their notoriously expensive tensor products, eSEN trails behind the completely non-equivariant Orb models in speed, at least in theory. Not only due to its conservative architecture (they also provide a direct-force version) but primarily because every message passing step requires a custom Wigner-D matrix for rotations into and out of the local frame of reference, a costly operation.
Orb-v3 (2025)
The Orb-v3 models from Rhodes et al.
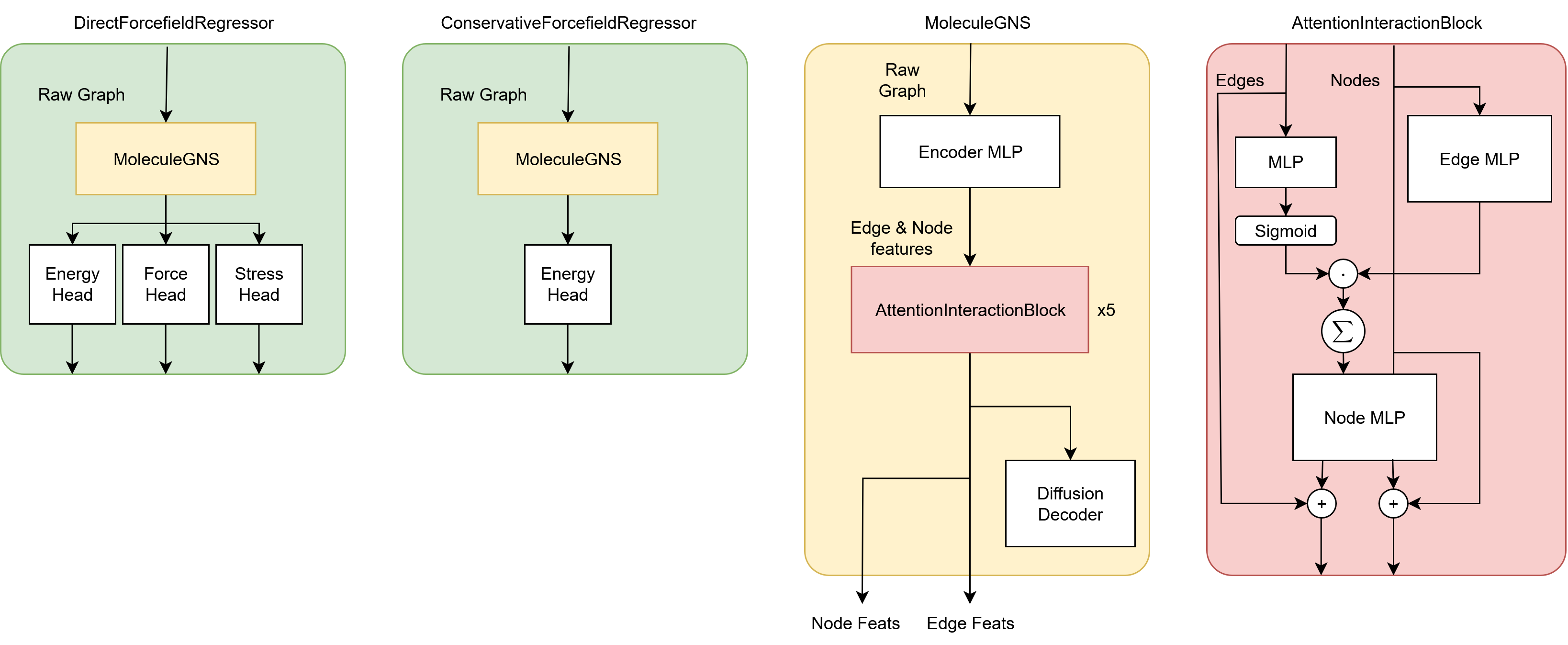
They provide both a conservative and direct-force version of the model, just like they did for Orb-v2. The explicit, direct-force version of Orb-v3 is distilled from the implicit, conservative model
generator = torch.zeros_like(cell, requires_grad=True)
rotation = rotation_from_generator(generator)
positions = (rotation @ positions[:, :, None])[..., 0]
inputs = [positions, displacement, generator]
grads = torch.autograd.grad(
outputs=[energy], # (n_graphs,)
inputs=inputs, # (n_nodes, 3)
grad_outputs=[torch.ones_like(energy)],
# ...
)
rotational_grad = grads[2]
rotational_grad_rms = torch.linalg.norm(
rotational_grad,
dim=(1, 2),
).mean()
loss += rotational_grad_rms * rot_grad_weight
Additionally, they include angular spherical harmonic embeddings as features and use 8 Bessel bases instead of the Gaussian RBF expansion
UMA (2025)
The UMA models from Wood et al.
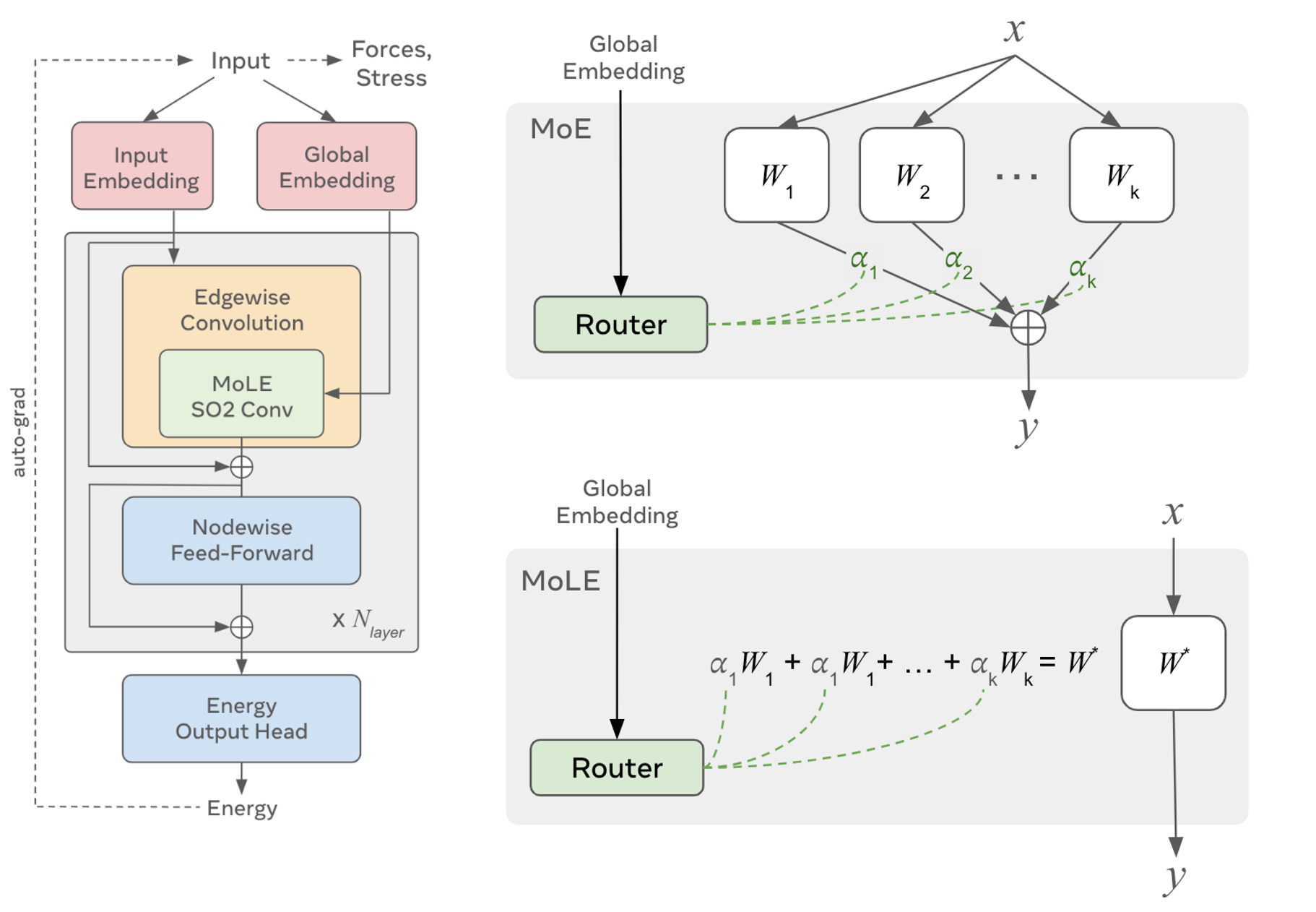
They introduce MoLE layers, essentially a mixture of experts
A short summary of the different MLIP models is shown in the table below,
| Model | Implicit | Equivariance | A.P. | N.L. | Cutoff |
|---|---|---|---|---|---|
| Orb-v2 | × | × | 25.2M | 20 | 6 Å |
| Orb-v3 Direct | × | × | 25.5M | 120 | 6 Å |
| Orb-v3 Conservative | ✓ | × | 25.5M | -- | 6 Å |
| Nequix | ✓ | e3nn | 708k | -- | 6 Å |
| MatterSim v1 | ✓ | e3nn | 4.5M | 256 | 5 Å |
| SevenNet MF-ompa | ✓ | e3nn | 25.2M | -- | 6 Å |
| eSEN | ✓ | SO(2) conv | 30.2M | -- | 6 Å |
| UMA M | ✓ | SO(2) conv | 50.0M | -- | 6 Å |
MD Frameworks
ASE
The Atomic Simulation Environment (ASE) is a molecular dynamics (MD) framework tool-set for Python with a strong focus on simplicity and ease-of-usease.Atoms object containing the atomic positions, species, cell vectors and boundary conditions of the system, and outputs the computed potential energy and forces acting on each atom. Optionally, they can choose to calculate other properties like stress for NPT simulations or charges for electrostatic interactions. This allows ASE to integrate potentials and dynamics from the vast majority of MD frameworks, simply by implementing this interface
Integrating MLIPs into ASE
The overwhelming majority of MLIP models provide a ready-to-use interface for ASE in the form of a calculatorcalculate method and fetches the computed properties as required.
LAMMPS
LAMMPS (Large-scale Atomic/Molecular Massively Parallel Simulator) is a highly scalable and parallelizable MD simulation framework leveraging empirical and classical potentials
Packages. A key strength of LAMMPS is its matured ecosystem of around 60 different packagesKSPACE with a long range electrostatic solver like particle-particle particle-mesh (PPPM)KOKKOS package for compilation of established styles
Integrating MLIPs into LAMMPS
As of writing this blog post, integrating MLIP models into LAMMPS is still a complicated and tedious task. There are two big roadblocks one will encounter for all integration pathways:
Programming language gap.
Even though the overwhelming majority of LAMMPS is written in C++, the overwhelming majority of ML models are written in Python, and not all of them use the same framework: Some researchers use PyTorchtorch.jit.compile, jax.jit or by herself; or she creates a bridge between the two languages using Cython and Numpy as C++ array interface for Python code.
The first option, Python to C++ compilation, is definitely faster and easier to use, provided it does actually compile. For example, when trying to port the Orb model*kwargs arguments and Optional[] types, which are non-trivial to remove in a large codebase in a limited time-frame. Also, this particular model family makes use of an adaptive graph construction scheme, which also seems to clash with TorchScript.
A C++-Python bridge is naturally slower, but framework-agnostic and significantly more versatile, given one has time and expertise to write the complicated cython interface, which is often non-trivial and must be done with the utmost care to avoid memory leaks and segmentation faults.
Different force philosophies.
LAMMPS is primarily built around the idea of pair styles or pair potentials
While this approach is mostly reasonable for most empirical potentials, MLIPs are designed to output the overall per-atom forces $F_{i}$ directly or as a gradient of the potential energy $U$ with respect to the atomic positions $r_{i}$: And that is not just a different design philosophy, it is the very reason why MLIPs are as accurate as they are. They should ideally operate holistically in the atoms’ neighborhood, \(F_{i} = f(\{r_{ij}, r_{ik}, \dots\}, \dots)\) instead of focussing just on pairwise interactions that fail to capture important many-body effects like angular dependencies (e.g. in H2O). Here, $j, k \in N(i)$ denotes all neighbors of atom $i$. To bridge this gap, two options make themselves available: In the philosophy of GDML from Chmiela et al.C++ and LAMMPS’s internal workings.
Option 1: New LAMMPS package
The most complicated, time-consuming and redundant way of integrating MLIPs into LAMMPS is the creation of a new package that provides a custom pair_style. This would require writing a new pair_style class in C++ that interfaces with our MLIP models, either via C++-Python bridge or by porting the model to C++ as discussed earlier in the programming language gap section.
While this is by far the most flexible option with the highest level of control, it requires the most interation with LAMMPS’s core codebase, is the most time-consuming and might need additional maintenance in the future. Popular models like MACE
Option 2: The ML-IAP package
To provide a universal interface for MLIP models, the ML-IAP package was added to LAMMPS in 2020. It uses a cython C++ to Python bridge to interface with Python code, allowing for on-the-fly integration of models without the need for a port. To integrate a brand-new MLIP model into LAMMPS when acting through python, one can either specify a loaded python module inheriting from the abstract MLIAPUnifiedInterface class performing the model invocation, or specify a .pt file containing such conforming model serialized by torch.save. This means that at the time of writing, one can not load a JAX model from a file, although support for it is underway
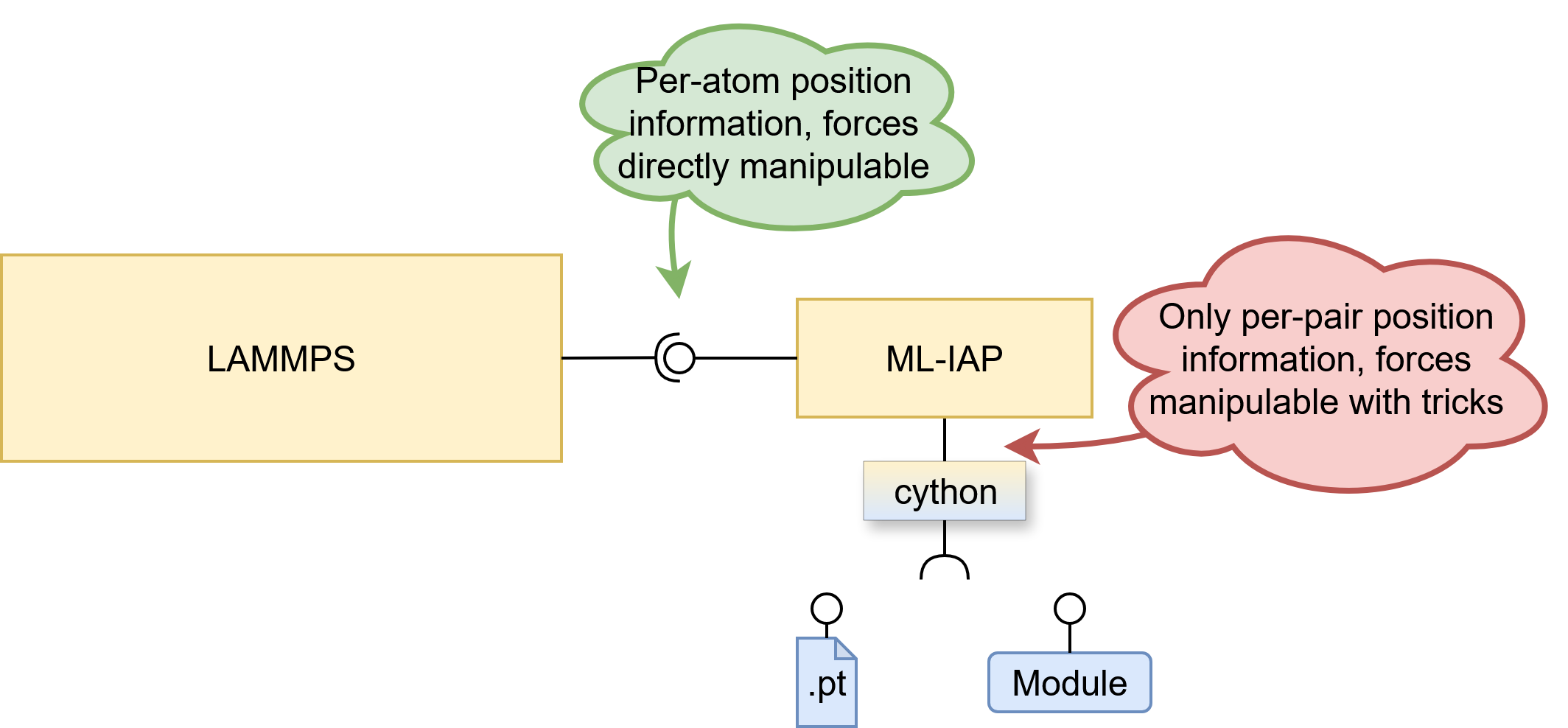
A big shortcoming of the provided cython bridge is the absence of absolute atomic positions. One is just given the pairwise distance vectors, which are often not enough. Not only do almost all models expect to receive absolute atomic positions, some model families like OrbML-IAP package typically expects the model to output the pairwise forces $F_{ij}$ instead of the overall per-atom forces $F_{i}$, as discussed in the different force philosophies section above. This means that one either folds and rewrites the model to conformity like NequIP
Option 3: Metatomic
In search for integration alternatives, the recent MetatomicMetatomic, it can be used seamlessly in all supported frameworks without any additional work.
Discussing the Options
Option 3 would have been the most elegant, future-proof and helpful solution. so it was naturally considered first. The primary downside is a C++ porting requirement with torch.jit.compile, neither always possible for torch models, nor applicable to JAX models like NequixML-IAP package would turn out unpatchable, which left us only with option 2, the ML-IAP package.
Modifying LAMMPS
ML-IAP already had access to the atoms->x array naturally, but didn’t provide it to the cython interface. To give the python code access, a pointer to the position array atom->x was added to the MLIAPData class, which is then wrapped by a cython interface class MLIAPDataPy which one can then use in python. For an efficient and zero-copy transfer of the C++ array to python, numpyatoms->ntotal exists, but this number varies over time, even with the same number of atoms as input, due to – which took some figuring out – the presence of ghost atoms. LAMMPS uses so-called ghost atoms for inter-worker communication, allowing the simulation of larger systems via domain decomposition as described by Plimpton
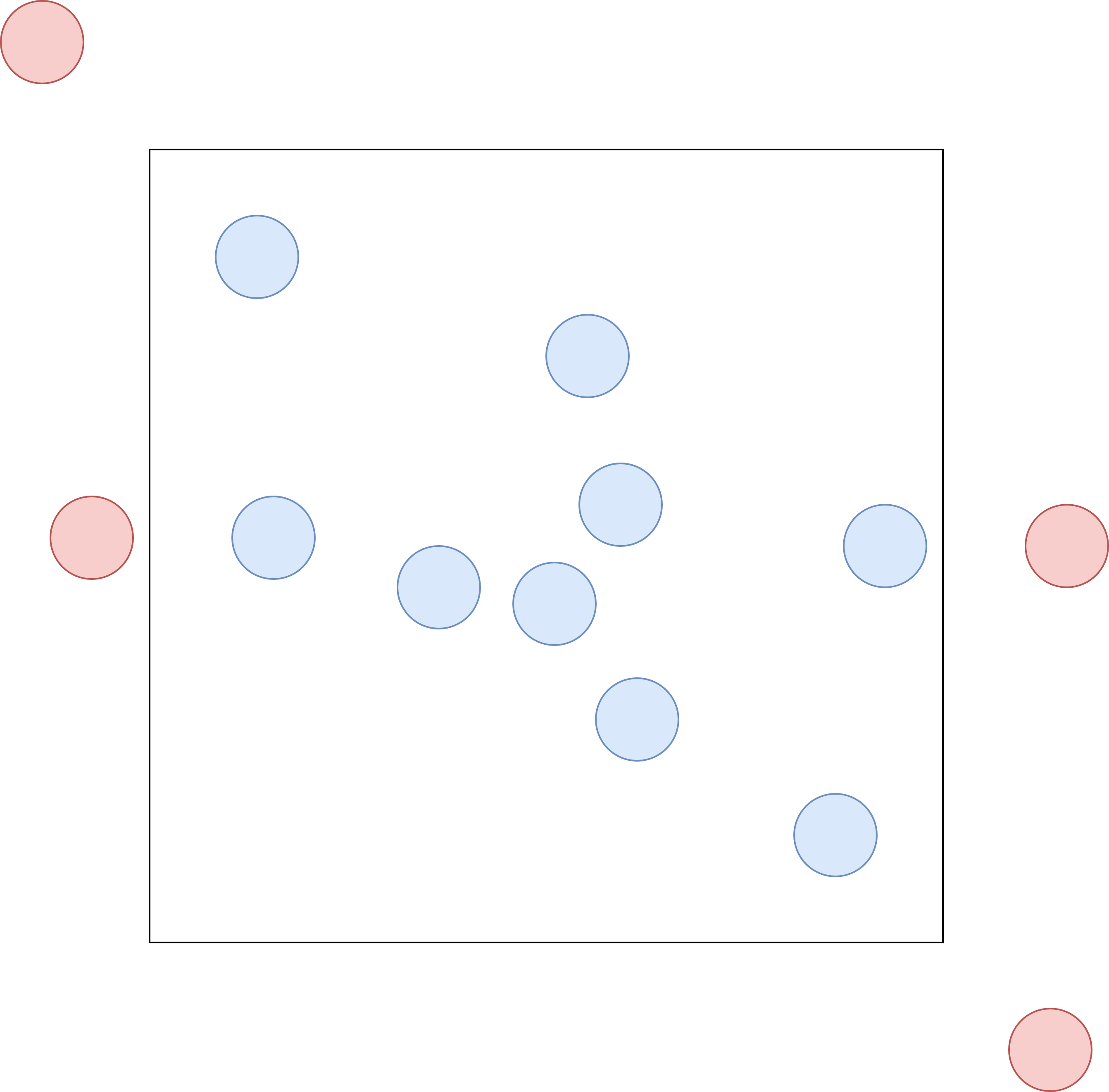
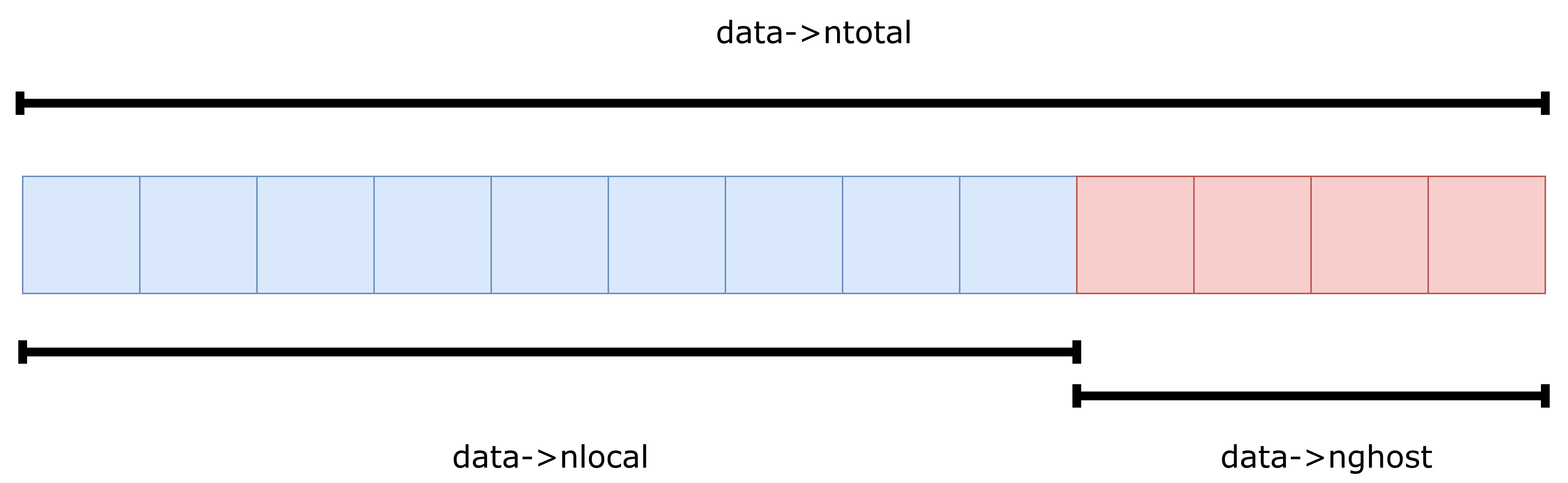
so it wasn’t exactly clear what type of atom can be found at what position in the atoms->x array. After some digging through the LAMMPS source code and some trial-and-error, two facts became clear: Both atom->x and atom->f contain the positions and forces of all atoms, local and ghost. Secondly, the arrays start with local atoms, directly followed by all ghost atoms as illustrated in the memory-layout panel above. We use all atoms, local and ghost as input to our MLIPs, because ghost atoms too can exert forces on real atoms as shown in the ghost-atom schematic above. We only apply the predicted forces to our local atoms, and postulate that if there exists a ghost atom, it is updated by exactly the same force predictor. This is important to ensure that Newton’s third law holds, i.e. that every action has an equal and opposite reaction. As was learned only later, LAMMPS uses ghost atoms not only for inter-worker communication, but also to simulate periodic boundary conditions, just like in the earlier illustration. Models are informed that the system is non-periodic and the rest is handled by LAMMPS automatically. This also means that models that previously didn’t support periodic boundary conditions, now do. Additionally, if one is not interested in the stress tensor, the original cell can simply be replaced by a large cubic cell, which we did for simplicity and to avoid any potential issues with out-of-cell atoms. We recognize that this solution requires the MLIP to compute more interactions than strictly necessary, resulting in an overhead of approximately 20-30%. Future work could focus on adding support for periodic boundary conditions naturally, ignoring ghost atoms. For this purpose, cell and boundary information has also been made available in the MLIAPData class.
All roads lead to ASE
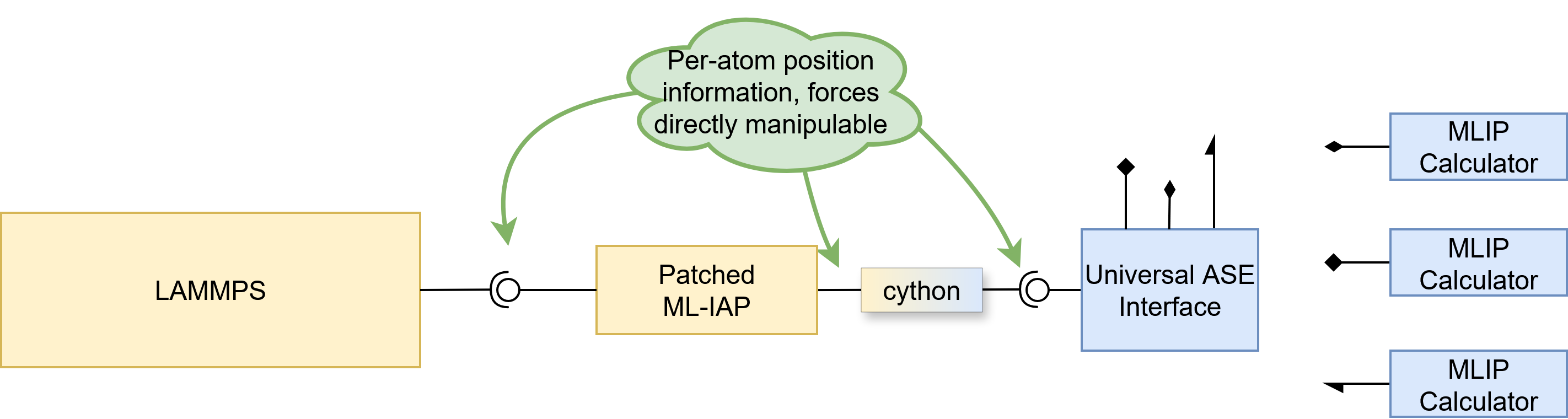
Pretty much all state-of-the-art (SOTA) models provide an ASE calculator interface, which has become the de-facto standard. There is no reason to waste time and effort writing complicated integrations for each and every model, when one can instead just write a small UniversalASEUnifiedMLIAPInterface class that combines both worlds seamlessly. Note that only the real and metal LAMMPS units are supported. ASE uses metal units, eV for energies and eV/Å for forces, while LAMMPS real units use kcal/mol for energies and kcal/(mol·Å) for forces, which we automatically convert using the conversion factor 1 eV = 23.0621 kcal/mol
Evaluation
Inference Speed
We measure inference speeds of force prediction as primary MD task on homogenous uniformly sampled systems of up 100,000 Helium atoms with an average density $\rho$ of 0.033 $\text{atoms/Å}^3$. To reduce variance of measurements caused by startup tasks like JIT compilation, a 40-step warm-up is performed on a system of 5 atoms. Then, we measure the inference speed of MLIPs on systems of 10, 100, 1.000, 10.000, and 100.000 atoms. We repeat the measurements a total of 20 times for each system size and model, and discard the first measurement to reduce size-depended startup tasks found in models like Orb-v3torch.compile through ASE calculator options and for JAX models using JAX’s jit compilation.
All models exhibit linear scaling with respect to the system size, meaning that MLIPs have the same computational complexity class as empirical force fields. Simulating a system twice as large simply requires twice the resources or double the time, which is not the case for any ab-initio method and a major advantage of MLIPs. However, the fact that all models scale linearly on the CPU and GPU does not mean that they are equally fast in practice. Using a GPU for inference results in speedups two orders of magnitude, as long as the system is larger than about 1.000 atoms. Inference times for smaller systems are bottlenecked by the general overhead and data transfer to and from the GPU. So for applications like high-throughput screening batching smaller systems to a single, clustered system can improve throughput by a large margin. Even though a single MD run can’t be batched due to its sequential nature, independent parallel-running MD simulations of smaller systems can be batched together and benefit. But not only the GPU has a large effect on the linear factor: Orb-v3 Direct predicts forces of 100.000 atoms as fast as UMA can predict the forces of only about 2.000 atoms. So even though in theory all models scale linearly with system size, O(N); the actual inference speed is model dependent. Explicit models without an equivariant architecture like Orb-v2 and direct Orb-v3 run the fastest. Even with more parameters than Orb-v2 and a higher neighbor limit, Orb-v3 ends up being slightly faster due to reduced overhead of edge aggregation and message passing of 10 layers fewer
In the following table we converted the raw inference speed into a more intuitive unit of nanoseconds per day (ns/day), assuming that a simulation is run with a time step of 1 fs ($10^{-15}$ s). This is a more common unit to express simulation speeds in the MD community, as it intuitively relates to how much simulation time can be covered in a day of wall-clock time.
| Model | 10 atoms | 100 atoms (↓) | 1,000 atoms | 10,000 atoms | 100,000 atoms |
|---|---|---|---|---|---|
| Orb-v3 Direct | 16.14 | 15.55 | 8.95 | 1.29 | 0.08 |
| Orb-v2 | 11.57 | 11.04 | 6.79 | 0.92 | 0.06 |
| Nequix | 12.00 | 8.91 | 2.41 | 0.25 | -- |
| Orb-v3 Conservative | 7.46 | 7.00 | 3.15 | 0.36 | -- |
| MatterSim v1 | 5.40 | 5.10 | 2.84 | 0.41 | -- |
| SevenNet MF-ompa | 1.82 | 1.51 | 0.51 | -- | -- |
| eSEN OAM | 1.40 | 1.19 | 0.24 | -- | -- |
| UMA M | 0.78 | 0.84 | 0.16 | 0.01 | -- |
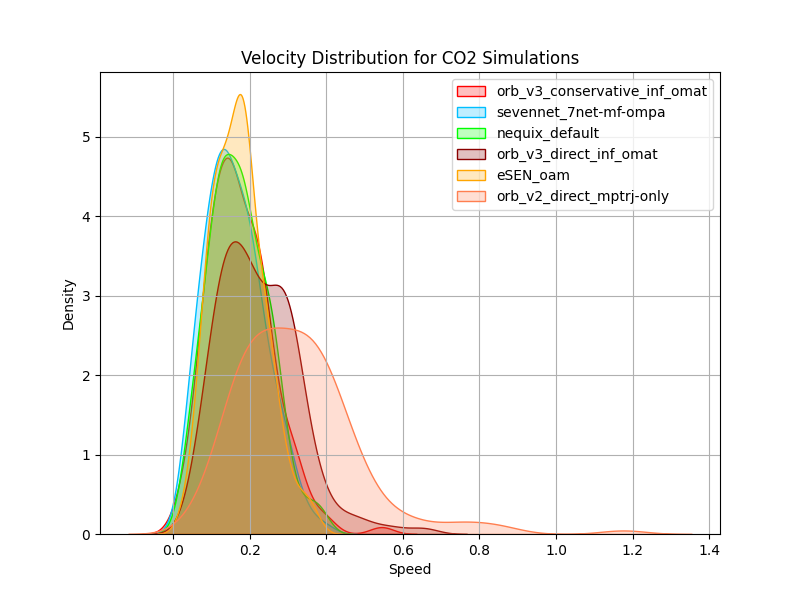
Simulating CO₂ Diffusion
We want to investigate how well MLIPs can run carbon-dioxide gas simulations in a NVE ensemble depending on temperature. For this we both investigate energy conservation and leverage diffusion coefficients as a measure of how well the MLIP captures the dynamics of CO₂ gas. The total energy of a system should be conserved in a NVE ensemble by definition. In MD simulations, the potential energy $E_{\text{pot}}$ of the system is converted to kinetic energy $E_{\text{kin}}$ and vice versa, keeping the total energy $E_{\text{tot}} = E_{\text{pot}} + E_{\text{kin}}$ constant over time. We evaluate energy conservation by running a system of 50 carbon-dioxide molecules (150 atoms) in a 25 Å × 25 Å × 25 Å periodic box. The system is first thermalized to 300K by sampling atomic velocities from a Maxwell-Boltzmann distribution and running a NVT ensemble for 10ps with a time step of 1 fs. Then, we re-sample the atomic velocities again from the Maxwell-Boltzmann distribution at 400K to introduce a temperature jump, and run a NVE ensemble for 10ps again. We repeat this procedure a total of 10 times and keep track of all relevant parameters during the simulation. Additionally, we calculate and compare the diffusion coefficients obtained from the MD simulations to the experimentally obtained diffusion coefficients of CO₂ gas at different temperatures
which we approximate. A high diffusion coefficient means that particles are moving quickly and spreading out rapidly, while a low diffusion coefficient means that particles are moving slowly and spreading out more slowly. Capturing this behavior correctly is a requirement for later simulating gas adsorption in MOFs, so we want to make sure that our models can do this correctly. The relationship between temperature $T$, collision frequency $\Omega(T)$, and diffusion coefficient $D$ in gasses is
\[D \propto \frac{T^{3/2}}{p\Omega(T)}\]where $p$ is the pressure of the gas in bar
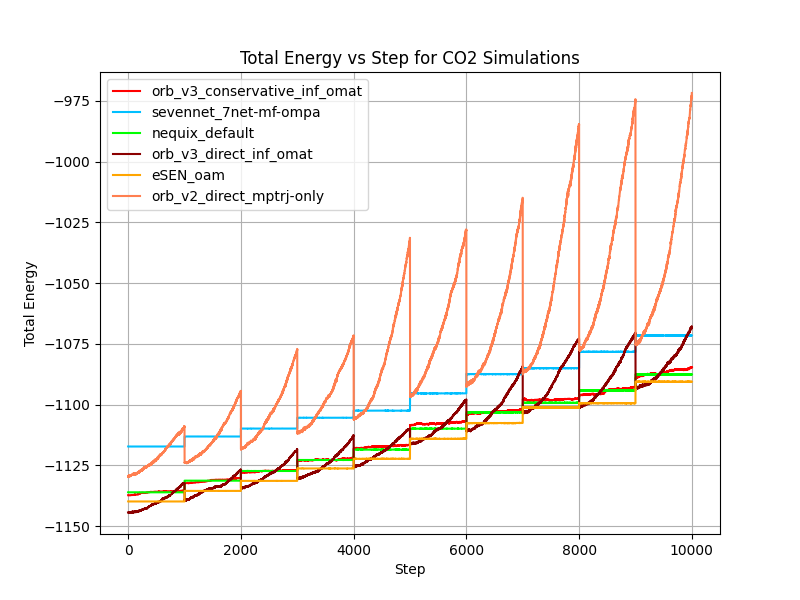
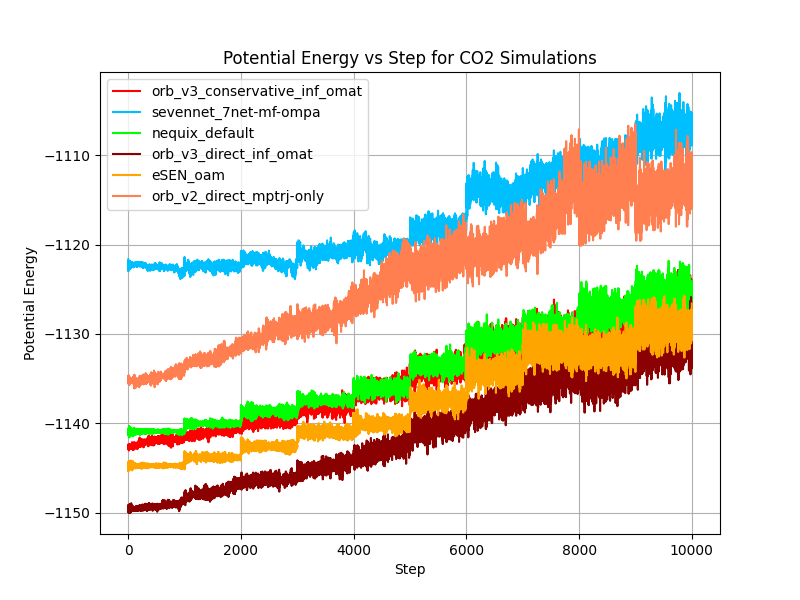
In the figure above we show the total and potential energy over time for different models during the CO₂ gas stability test. Orb-v2 and direct Orb-v3 exhibit significant energy drift over time, 241.52 meV/ps and 78.14 meV/ps on average, respectively. The argument can be made that this is a consequence of the explicit architecture, and we agree to some extent. Learning a smooth energy surface is arguably easier than learning a smooth force surface, because it allows the model to focus on the relationship between atoms rather than figuring out how this relationship translates into forces, where it has to learn what a force is and how it rotates with the system, etc. But the conservative Orb-v3 model also experiences a slight energy drift of 9.42 meV/ps on average, despite being “conservative” by design.
| Model | Drift (meV/ps) |
|---|---|
| Orb-v2 | 241.52 |
| Orb-v3 Direct Inf | 78.14 |
| Orb-v3 Conservative Inf | 9.42 |
| SevenNet MF-ompa | 0.30 |
| Nequix Default | 0.26 |
| eSEN-30M-OAM | 0.26 |
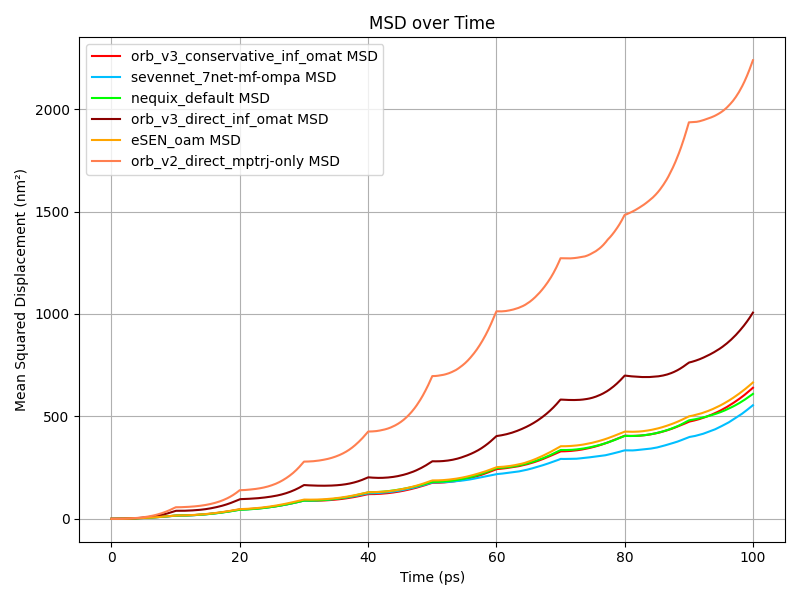
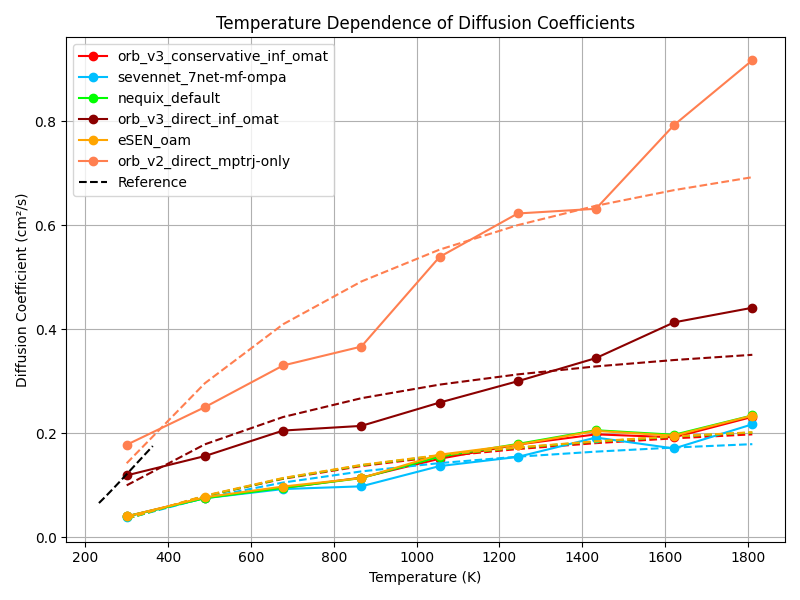
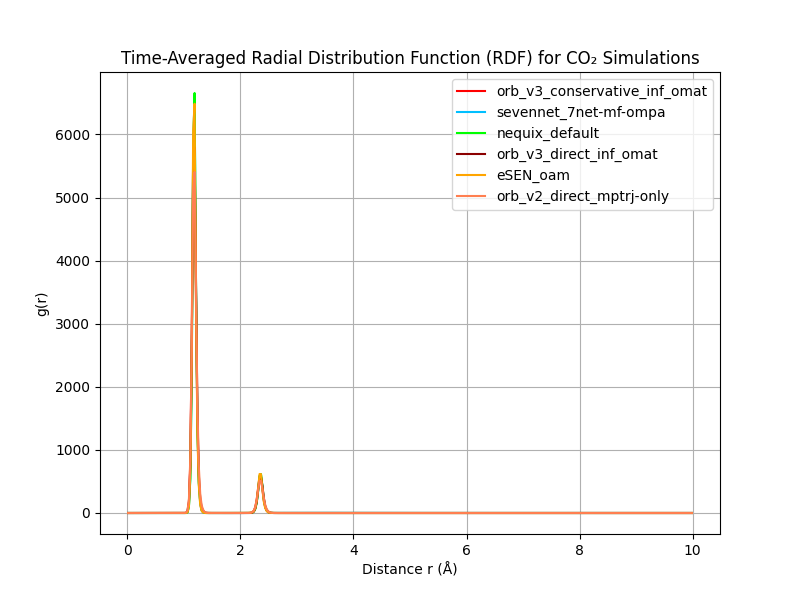
In the MSD figure above we show the MSD of CO₂ molecules over time for different models (left) and use that to compute the diffusion coefficients at different temperatures (right).
They show Orb-v2 and direct Orb-v3 exhibiting a diffusion coefficient twice and thrice as large as all other models. However, Burgess
We fit parameters $A$ and $B$ to
\[\ln(D) = A + \frac{B}{T}\]a simple Arrhenius-type equation
| Model | $D_{\text{ref}}$ / cm²s⁻¹ | $A$ | $B$ |
|---|---|---|---|
| Experimental | 0.120 | 0.015 | -637.089 |
| Orb-v2 | 0.178 | 0.0533 | -568.4568 |
| Orb-v3 Direct | 0.191 | -0.7975 | -451.4263 |
| Orb-v3 Conservative | 0.004 | -1.2809 | -612.5149 |
| eSEN-30M-OAM | 0.004 | -1.2570 | -618.9303 |
Simulating CO₂ Adsorption in MOF
Metal-Organic Frameworks (MOFs) have become highly popular among materials scientists, primarily due to their exceptional porosity and tunable properties
CO$_2$ is the primary greenhouse gas responsible for climate change
Electrostatic interactions are fundamental forces between charged particles and play a critical role in a wide range of physical, chemical, and biological processes. These forces can be modelled with Coulomb’s law
where $q_{1}$ and $q_{2}$ are the charges of the interacting particles, $r$ is their separation distance, and $\epsilon_{0}$ is the permittivity of free space. The $1/r$ dependence of electrostatic interactions makes them exceptionally long-ranged, meaning they cannot be neglected even at distances exceeding 25 Å. In our MOF example, the interaction energy between the $Mg^{2+}$ metal node ($q\approx1.45e$
Van-der-Waals interactions are the other primary force acting on CO$_2$ molecules in Mg-MOF-74, and as discussed in the Kohn-Sham section they require dispersion corrections to be modelled accurately in DFT simulations. The effects of VdW forces fall off with $1/r^{6}$ and are therefore rather short-ranged compared to electrostatic interactions. While some models like Orb-v3SumCalculator object that simply sums the energies and forces of both calculators. Contrasting the need for a post-processing correction, the UMA model familyOMol25
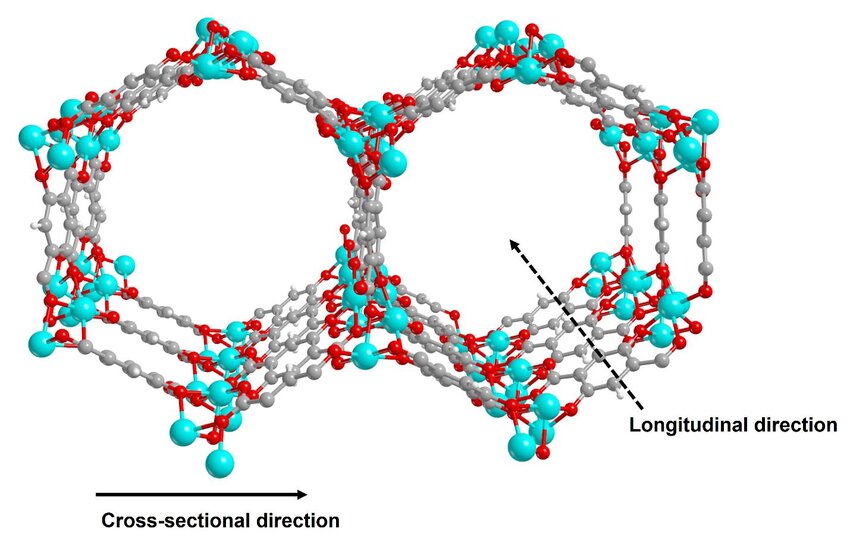
To benchmark MLIPs in a realistic setup, we simulate the adsorption of CO2 in Mg-MOF-74 via MD simulations. We closely follow and make use of the scripts provided by Fu et al.
| Simulation | 0 | 1 | 2 | 3 | 4 | 5 | 6 | 7 | 8 | 9 |
|---|---|---|---|---|---|---|---|---|---|---|
| Pressure (bar) | 0.000 | 0.073 | 0.151 | 0.237 | 0.330 | 0.433 | 0.547 | 0.677 | 0.825 | 1.000 |
| Pressure (kPa) | 0.000 | 7.3 | 15.1 | 23.7 | 33.0 | 43.3 | 54.7 | 67.7 | 82.5 | 100.0 |
For simplicity, we convert 1 bar to 100 kPa instead of 101.325 kPa during post-processing. Every 10 simulation steps, we dump the entire system state to a lammpstr file, about 8 GB per simulation. To compress the output data, we convert the lammpstr files to a custom delta-compressed binary format with gzip compression, resulting in a trajectory file of about 100MB per simulation. To compute the equilibrium adsorption capacity, we only consider the last 50\% of each simulation, and average the number of adsorbed CO2 molecules over time.
Unfortunately, the adsorption isotherm calculation could only be performed using the Orb-v3 direct-force model and the original empirical potentials from Fu et al.
Running the fast direct-force Orb-v3 model on the LAMMPS-extended Mg-MOF-74 carbon-dioxide filled system
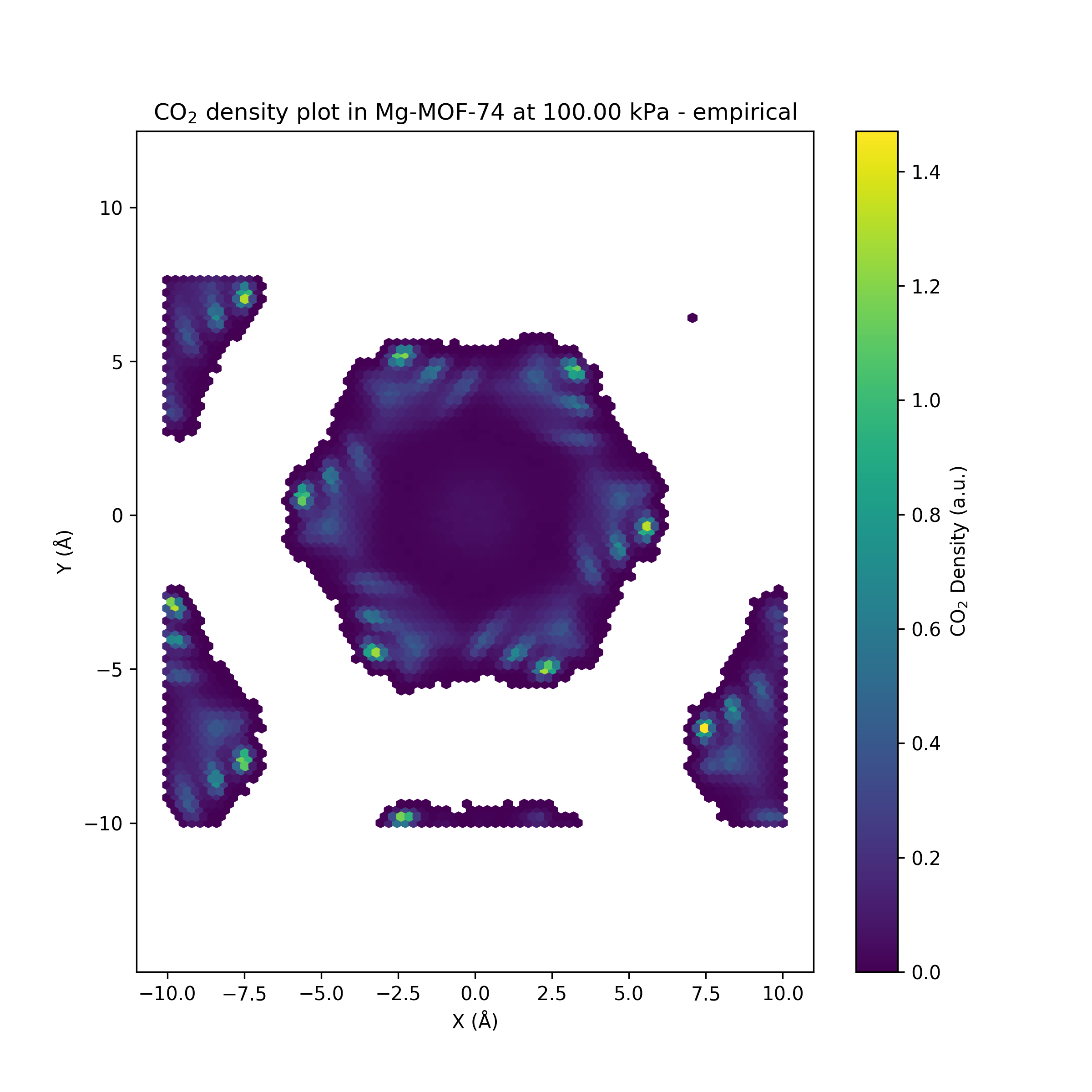
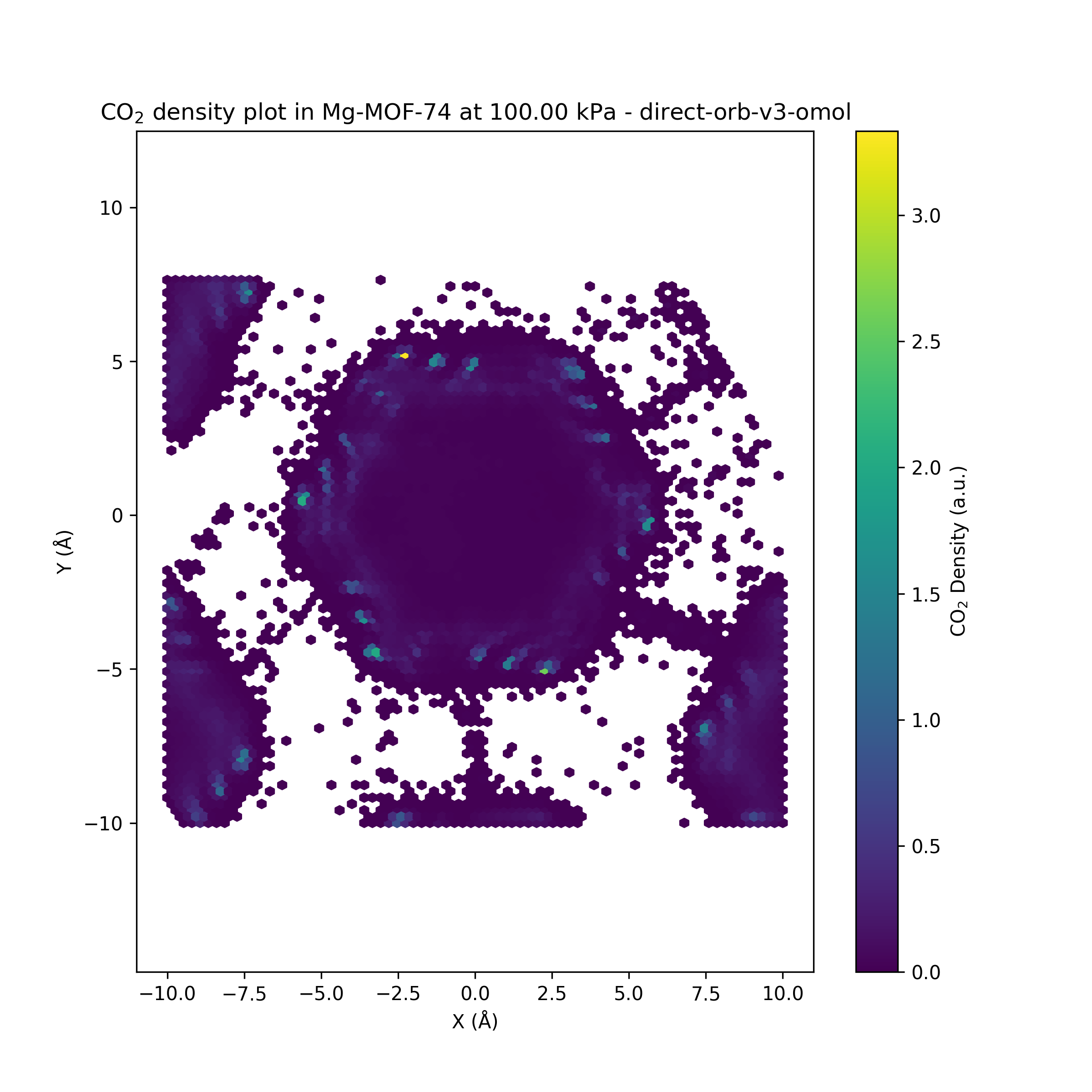
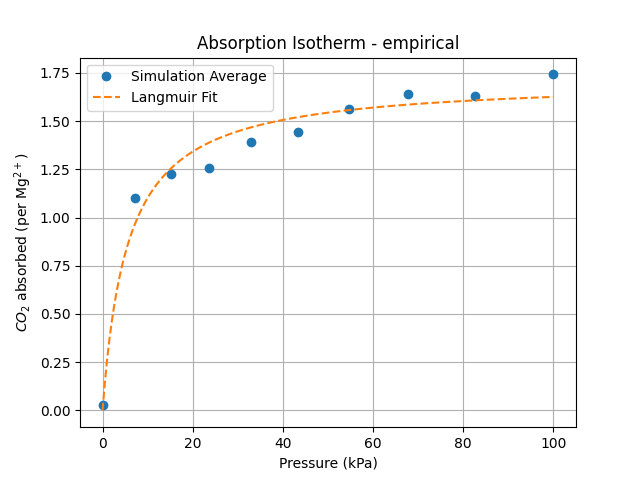
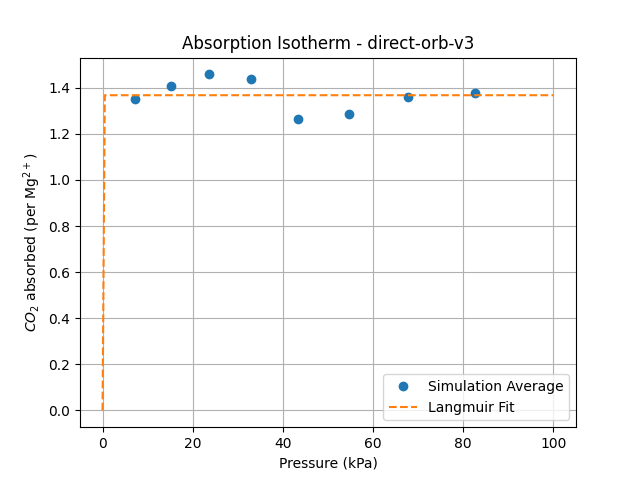
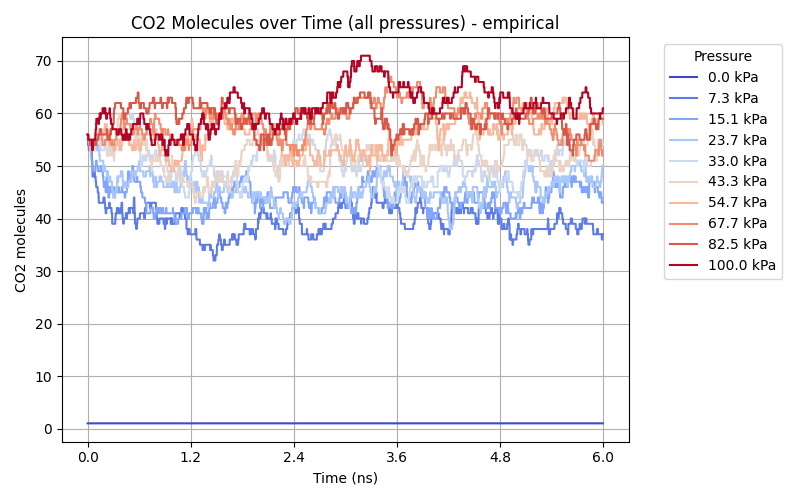
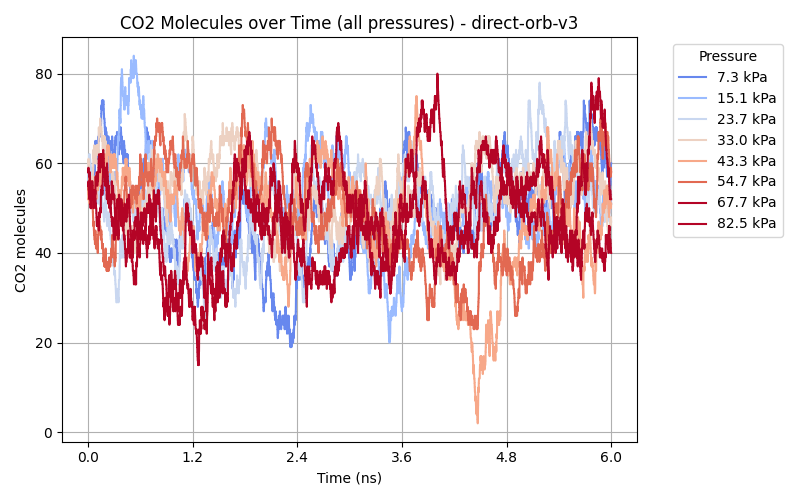
While the Orb-v3 direct-force omol model can reproduce some of the results from
to both isotherms, where $q$ is the amount adsorbed at pressure $P$, $q_{\text{max}}$ is the maximum adsorption capacity, and $b$ is a constant related to the affinity of the adsorbent for the adsorbate. This model is plotted as a dashed line in the adsorption isotherm figure, showing a good fit for the empirical results while the Orb-v3 direct-force omol model predicts almost no pressure-related correlation for CO$_2$ adsorption. Finally, we plot the number of CO$_2$ molecules inside the GCMC simulation over time for different pressure settings in the figure above. While the empirical model shows a small variation in the number of CO$_2$ molecules over unique pressure-dependent mean values over time, the results obtained from the Orb-v3 direct-force omol model seem random with a high variance without any pressure-dependent mean value. A potential area of future work could be the evaluation – with the necessary computational resources at hand – of more models or orb-v3 variations to find better performing models for this task.
The source code for all the experiments in this post is available on GitHub
Conclusion
In this post, we explored how MLIPs work and how they are built, different architectures and design choices that go into making them. We were successful in building a universal ASE-MLIP interface for LAMMPS, allowing us to run an originally empirical Mg-MOF-74 CO$_2$-adsorption GCMC simulation with an MLIP instead, only to discover that it doesn’t reproduce the more physically-sound empirical results. While we were able to evaluate the cheapest MLIP model on the CO$_2$ GCMC simulation, the more accurate models remained prohibitively expensive to run within a practical time frame. Since the field of MLIPs is still relatively in its early stages, we hope to see more exciting developments in the future. We specifically hope to see more work on evaluating MLIPs for predicting useful downstream physical and dynamical properties, and also on investing in stronger integration with established MD frameworks. New training schemes beyond single step energy prediction, that can efficiently incorporate these downstream physical properties is also an open question.
PLACEHOLDER FOR ACADEMIC ATTRIBUTION
BibTeX citation
PLACEHOLDER FOR BIBTEX